synthetic data will be added later and full example will be provided.
Description
This vignette describes the data structure required by the package
and demonstrates selected classification functions.
The healthpopR package uses a simple common data model
(CDM) with predefined variable names.
The expected datasets, variables, and their definitions are presented.
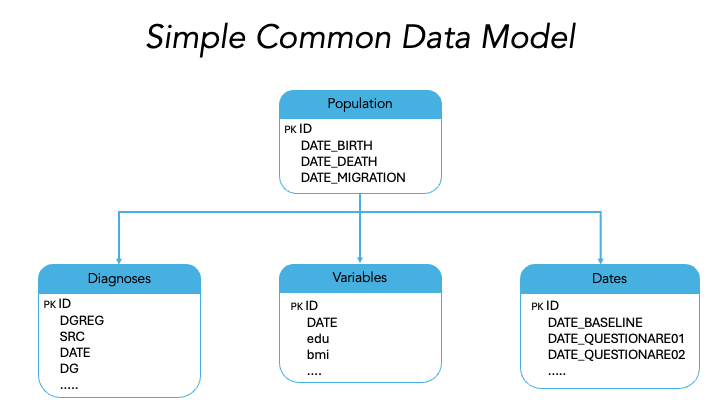
population
Population data should include only one row per ID and few DATE variables.
| Variable Name | Description |
|---|---|
| ID | Patient identifier |
| DATE_BIRTH | Date of patient birth |
| DATE_DEATH | Date of when patient died or NA |
| DATE_MIGRATION | Date of when patient migrated to abroad or NA |
diagnoses
Diagnoses data is follow-up of the patients diagnoses over the timeline.
| Variable Name | Description |
|---|---|
| ID | Patient identifier |
| DGREG | Diagnose Code Source (ICD10, ICD9 or ICD8) |
| SRC | Registry source (ex. hilmo, avohilmo etc.) |
| DATE | Date of the diagnose |
| DG | Full Diagnose Code (ex. E11, H903, H0410) |
| ICD10_3LETTERS | Only with ICD10 code. Diagnose Code in first 3 letters |
| ICD10_CLASS | Only with ICD10 code. ICD-10 Diagnose Class (Ex. E00–E90, H60–H95, H00–H59) |
| DATE_BIRTH | Day of patient birth |
| AGE | Only with murtumat Patient age on diagnose date |
other datasets
These will be modified in future. Now these are used only in cox modelling phases
Basic Data Handling
Load data
# synthetic raw datasets
population <- arrow::read_parquet("../data/synth_population.parquet")
diagnoses <- arrow::read_parquet("../data/synth_diagnoses.parquet")
population_variables <- arrow::read_parquet("../data/synth_population_variables.parquet")population data should have these variables:
| Variable Name | Description |
|---|---|
| ID | Patient identifier |
| DATE_BIRTH | Date of patient birth |
| DATE_DEATH | Date of when patient died or NA |
| DATE_MIGRATION | Date of when patient migrated to abroad or NA |
diagnoses data should have these variables:
| Variable Name | Description |
|---|---|
| ID | Patient identifier |
| DGREG | Diagnose Code Source (ICD10, ICD9 or ICD8) |
| SRC | Registry source (ex. hilmo, avohilmo etc.) |
| DATE | Date of the diagnose |
| DG | Full Diagnose Code (ex. E11, H903, H0410) |
To diagnoses data we will create also these variables:
ICD10_CLASS | Only with ICD10 code. ICD-10 Diagnose Class (Ex. E00–E90, H60–H95, H00–H59) |
DATE_BIRTH | Day of patient birth |
AGE | Only with murtumat Patient age on diagnose date |
population_variables should have these variables:
| Variable Name | Description |
|---|---|
| ID | Patient identifier |
| edu | Education, in our example it factor with categories Low, Med and High |
| bmi | Body Mass Index as double |
For bmi we will categorized variables.
Categorize ICD-10 Diagnoses
ICD-10 Codes are useful to categorize to 3 Letters and to upper category:
diagnoses <- diagnoses %>%
mutate(
ICD10_3LETTERS = healthpopR::recode_icd10_3letters(DG, DGREG = DGREG),
ICD10_CLASS = healthpopR::recode_icd10_class(ICD10_3LETTERS, DGREG = DGREG)
) %>%
left_join(population %>% select(ID, DATE_BIRTH), by = "ID") %>%
mutate(AGE = floor(lubridate::time_length(difftime(DATE, DATE_BIRTH), "years")))Categorize population_variables BMI
BMI can be categorized to 4 or 6 category classes. We can use these variables in Cox model in future.
population_variables <- population_variables %>%
mutate(
bmi_cat1 = factor(healthpopR::categorize_bmi(bmi, levels = 6)),
bmi_cat2 = factor(healthpopR::categorize_bmi(bmi, levels = 4)),
edu = factor(edu)
)